Study Timeline
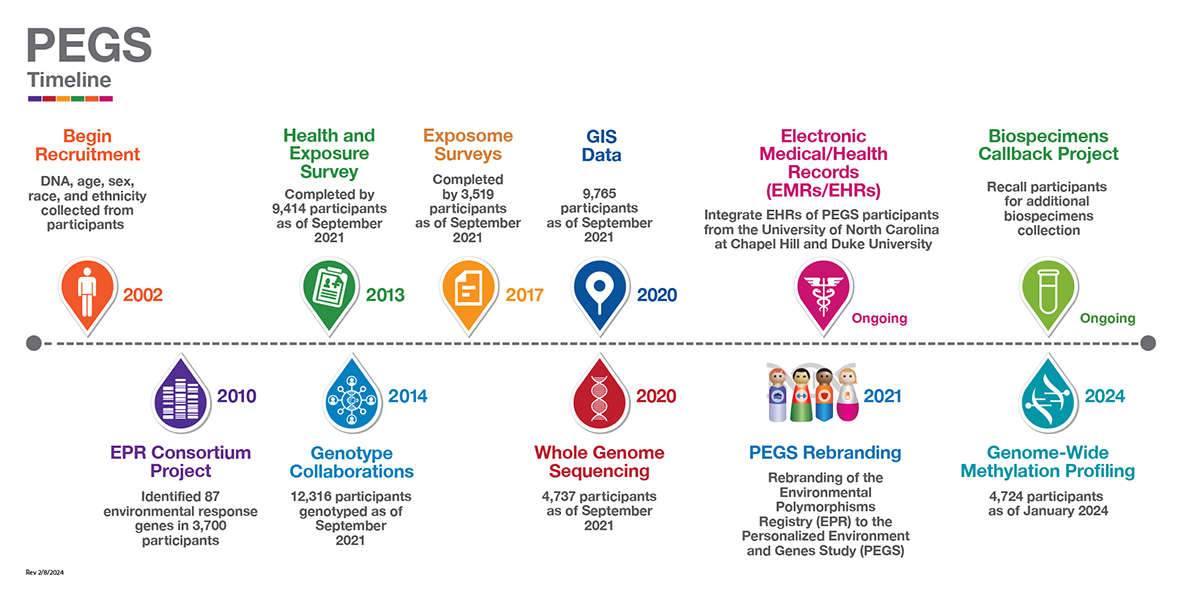
In 2002, Principal Investigators at NIEHS established the Environmental Polymorphisms Registry (EPR) as a core facility to recruit participants to meet ongoing research needs. EPR was a cross-sectional cohort with participants selected by convenience sampling. For the first ten years, the focus was on recruiting and enrolling participants through blood-draw laboratories at health care clinics, university campuses, health fairs, and drives at businesses, community centers and events. As part of clinical call-back studies, individual investigators at NIEHS completed the initial genotyping efforts. In 2010, the EPR Consortium Project was created as a pilot initiative to genotype 87 environmental response genes (656 single-nucleotide polymorphisms, or SNPs) in approximately 3,700 randomly selected participants.
While continuing to serve the important capacity of targeted and general recruiting, the scope of the cohort has expanded to include extensive data on pharmaceutical, lifestyle and environmental exposures, diseases and genetics. To reflect this focus, the EPR has now been renamed the Personalized Environment and Genes Study (PEGS).
Data Collection
Two surveys are used to collect phenotype and exposure data from participants.
Health & Exposure Survey
Beginning in 2013, participants were administered the Health & Exposure Survey to collect data on demographics, family medical history, lifestyle factors such as smoking and alcohol use, and occupational exposures.
Exposome Surveys
Beginning in 2017, participants were also administered the Exposome Survey to collect comprehensive information on endogenous and exogenous exposures throughout life.
Part A (External Exposome) – Part A asks about external exposures, including chemical and environmental exposures at work and home from childhood to the present.
Part B (Internal Exposome) – Part B asks about internal exposures, including medications, and lifestyle factors such as physical activity, stress, sleep, and diet.
Whole Genome Sequencing
Whole genome sequencing (WGS) enables the interrogation of common and rare variants and structural variations, including high-resolution human leukocyte antigen (HLA) variants. In 2019, the Broad Institute performed WGS for blood samples obtained from 4,737 PEGS participants with the most complete survey data. Quality control was performed for the WGS data, which were aligned to the hg38 human reference assembly to obtain single nucleotide variants and small insertions/deletions (indels). This resulted in approximately 43 million high-quality variants, which were annotated using the WGS annotator (WGSA). As part of this work, six-digit HLA genotypes and structural variants were identified.
Electronic Health Records (EHRs)
In 2019, linkage to electronic medical records (EMRs) from Duke University Health System and UNC Health at the University of North Carolina Chapel Hill was initiated. This linkage will provide an unprecedented integration of medical records across medical systems, exposure data, and WGS data. The electronic health records will enable the collection of long-term data on health and disease, in conjunction with multidimensional phenotypes that include laboratory data, images, vital signs, and other clinical information. This information can be used for understanding mechanisms and patterns of variability in disease susceptibility, disease evolution, and drug responses.
Geographic Information Systems (GIS) Data
In 2020, the addresses of PEGS participants were assigned mapping coordinates. This geocoding enabled proximity analysis of contaminant sources as surrogates for exposure and geospatial data linkages to obtain exposure estimates from publicly available data from federal and state regulatory agencies. For each participant, addresses from five study events were mapped: time of initial enrollment, time of completion of the Health and Exposure Survey, time of completion of the External Exposome Survey and the longest-lived childhood address and the longest-lived adult address from the External Exposome.
To obtain exposure estimates, the addresses are linked to Geographic Information System (GIS) databases, including:
- Center for Air, Climate, and Energy Solutions (CACES), which includes measures of air pollution such as carbon monoxide, nitrogen dioxide, ozone concentration, etc.
- North Carolina Department of Environmental Quality (NCDEQ), to calculate distance to various point sources such as swine Concentrated Animal Feeding Operations (CAFOs), hazardous waste site, hazardous spill site, EPA superfund site, wastewater treatment plant release site, etc.
- Toxics Release Inventory (TRI), which tracks the industrial management of toxic chemicals.
- Atmospheric Composition Analysis Group (ACAG), which estimates the surface area of ground-level particulate matter.
- Department of Transportation (DOT), Federal Aviation Administration (FAA), Federal Communications Commission (FCC) and the Nuclear Regulatory Commission (NRC) to calculate distances to major roadways and rail depots, airports, cell towers, nuclear power stations, etc.
- Modern Era Retrospective analysis for Research and Applications (MERRA-2) containing consistent estimates of climate and environmental metrics assimilated from a range of satellite-based environmental observations.
- CDC/ATSDR Social Vulnerability Index (SVI) to obtain the summaries of social determinants of health at the census tract level including an overall index, four component indexes (socioeconomic status, household characteristics, racial and ethnic minority status, and housing type/transportation), and source variables used to compute each index component (e.g., poverty, education, overcrowding, access to vehicle, etc.) available for 2010, 2014, 2016, and 2018.
Exposure estimates from additional GIS databases, with a focus on groundwater, natural disasters, weather, and temperature, will continue to be added to the growing data assembled on PEGS participants.
Participant Call-back for Additional Data Collection
The PEGS cohort is unique in the United States. Recent contact has been maintained with 6,053 participants, the majority (70%) of whom reside within 100 miles of the NIEHS Clinical Research Unit (CRU) based on their current address. The ability to recruit participants for call-back studies at the Clinical Research Unit has enormous potential for following up findings through additional data collection. Maintaining an adequate pool of participants who can be contacted for follow-up studies, requires ongoing recruitment through studies at the CRU, the NIH Clinical Center, the PEGS enrollment website, and Clinicaltrials.gov.
PEGS participants have consented to provide blood, cheek cells, exhaled breath, hair, household dust, nail clippings, nasal cells, saliva, skin cells, sperm, sputum, stool, baby teeth, urine and other tissues for specific studies. For participants with certain conditions of interest we may, with the consent of the participant, seek to obtain tissue samples from treating physicians and pathologists, collected during routine medical diagnosis and/or treatment, and routinely saved after biopsy or surgery. As needed, PEGS staff will work directly with participants and participants’ health care providers to obtain and complete any release forms required by the originating health care facility.



