DNA Repair & Nucleic Acid Enzymology Group
Protein and nucleic acid structures are currently being determined in the research community at a pace that was unimaginable until only recently. During 2004, an average of 15 structures were deposited per day at the Protein Data Bank. These structures potentially provide an in-depth understanding of the function of protein and nucleic acid molecules as well as their role in complex biological systems. However, while a structure can define the environment of an atom in molecular detail, functional studies are required to decipher these atomic interactions in the context of cellular events. Accordingly, both functional and structural analyses of proteins and nucleic acids offer tremendous promise for uncovering fundamental biological mechanisms. This sub-group’s broad research interest is to understand basic mechanisms of biological catalysis and recognition of substrates and inhibitors by relating catalytic efficiency or inhibitor potency to molecular structure through a combination of site-directed mutagenesis, kinetic analysis, molecular modeling and structure determination. An understanding of these basic mechanisms is fundamental for understanding catalysis in vivo, the rational design of novel inhibitors for therapeutic use, and the influence of the environment on cellular pathways.
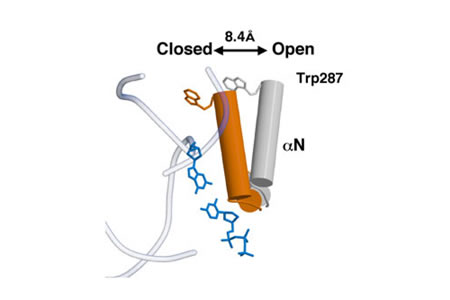
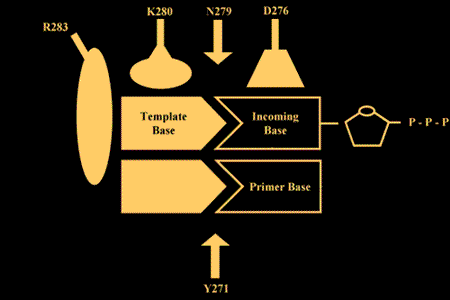
Relevant Reviews: Beard, W.A. and Wilson, S.H. Structure and mechanism of DNA polymerase β. Chemical Reviews, "DNA Damage and Repair," 106:361-382, 2006.
Researchers: William A. Beard, David D. Shock, Vinod Batra
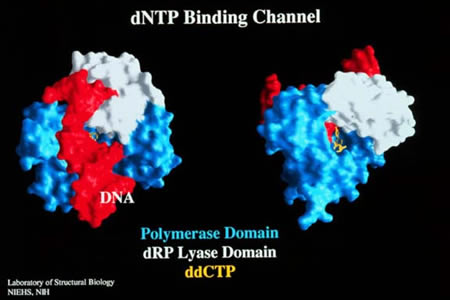
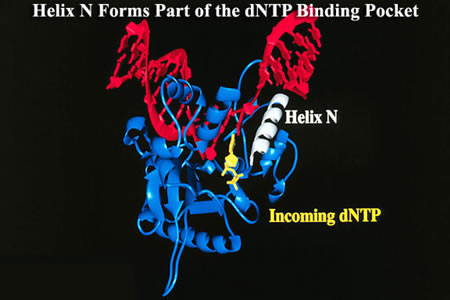
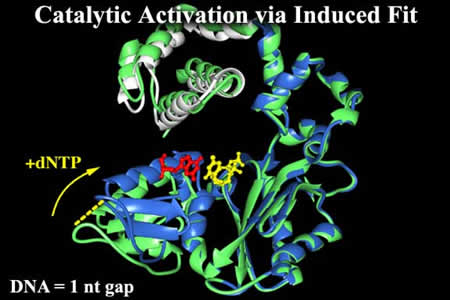
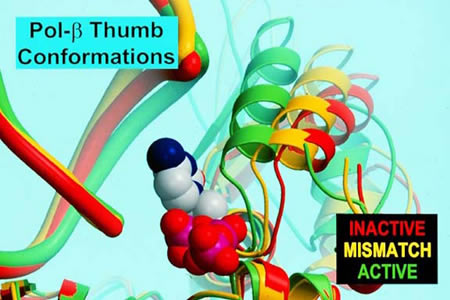
Slides