This package contains programs that fit log-linear models to estimate the relative risk associated with a candidate risk haplotype in a triad-based association study as described in the manuscript Shi M, Umbach DM, Weinberg CR "Using Case-parent Triads to Estimate Relative Risk Associated with a Candidate Haplotype." The supplementary results are available as the document "Comparison-to-Carayol-et-al.pdf."
TRIMMEST (69KB) is the ZIPPED file containing 8 items.
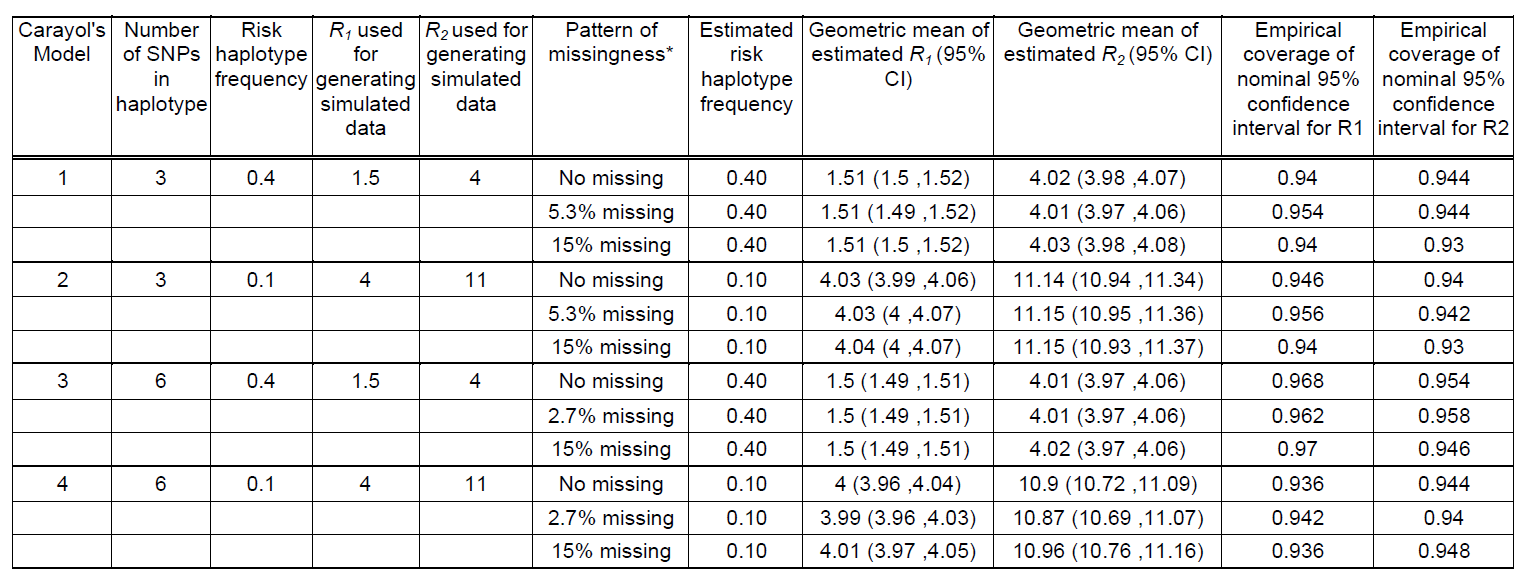
Table 1. Performance of log-linear model under Carayol et al's (2006) simulation scenarios (Table I of Carayol et al 2006) using 500 replicates of 1000 triads.
*This column represents the percentage of genotypes missing randomly, which is different from the missingness defined by Carayol et al. (the probability for an individual to have at least one unobserved SNP in a haplotype). Our scenarios of 5.3% and 15% missing genotypes correspond to their 15% and 39% missingness respectively.
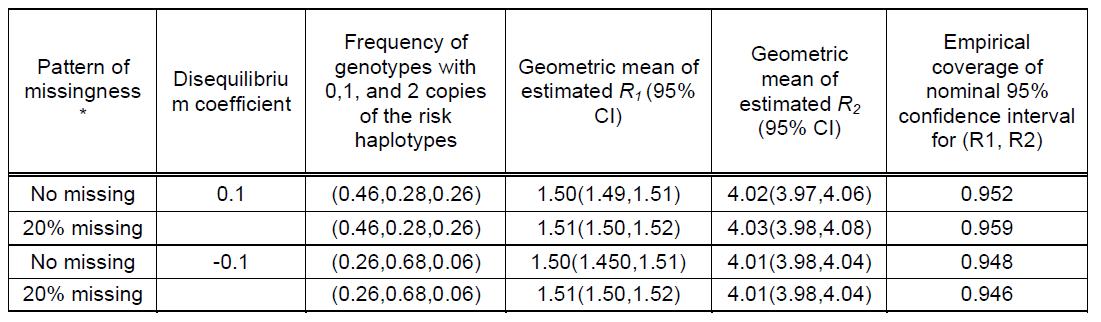
Table 2. Performance of log-linear model under Carayol et al's (2006) Hardy-Weinberg disequilibrium scenario (Table VIII of Carayol et al 2006) using 500 replicates of 1000 triads.
Note: Simulation conditions is Model 1 of Carayol et al. A total of 6 haplotypes comprised of 3 SNPs; risk haplotype frequency=0.4 * Our 20% missing genotypes correspond to Carayol et al's 49% missingness.
TRIMMEST Tables CSV version
Reference:
Carayol J, Philippi A, Tores F. 2006. Estimating haplotype relative risks in complex disease from unphased SNPs data in families using a likelihood adjusted for ascertainment. Genet Epidemiol 30(8):666-76.