Structure Function Group
SMCHD1 (structural maintenance of chromosome flexible hinge domain containing 1) is a 2005 amino acid protein involved in gene silencing, with reported roles in X-inactivation, genomic imprinting, and non-homologous end joining (NHEJ). Missense mutations in SMCHD1 are associated with arhinia, and Facioscapulohumeral Muscular Dystrophy Type 2 (FSHD2). In support of Dr. Shaw at the NIEHS, we solved the X-ray crystal structure of the SMCHD1 ATPase module (aa 24-580) (1). The ATPase module is composed of three domains: a previously uncharacterized ubiquitin-like domain (UBL), an ATPase domain, and a transducer domain. We determined that, while both the ATPase and transducer domains are required for ATPase activity, the UBL domain is not. However, the UBL is required for dimerization, consistent with the domain-swap observed in the crystal structure dimer. While variants that lead to FSHD2 are scattered throughout the protein, those associated with arhinia are located only in the ATPase module. Many of the mutations in both arhinia and FSHD2 appear to cluster at domain interfaces, suggesting they may affect conformational dynamics. Interestingly, of the disease associated variants tested, both arrhinia- and FSHD2-associated variants displayed some ATPase activity. Yet the FSHD2-associated variants displayed more severely diminished dimerization compared to the arrhinia-associated variants. We are continuing to study how these mutations may contribute to disease.
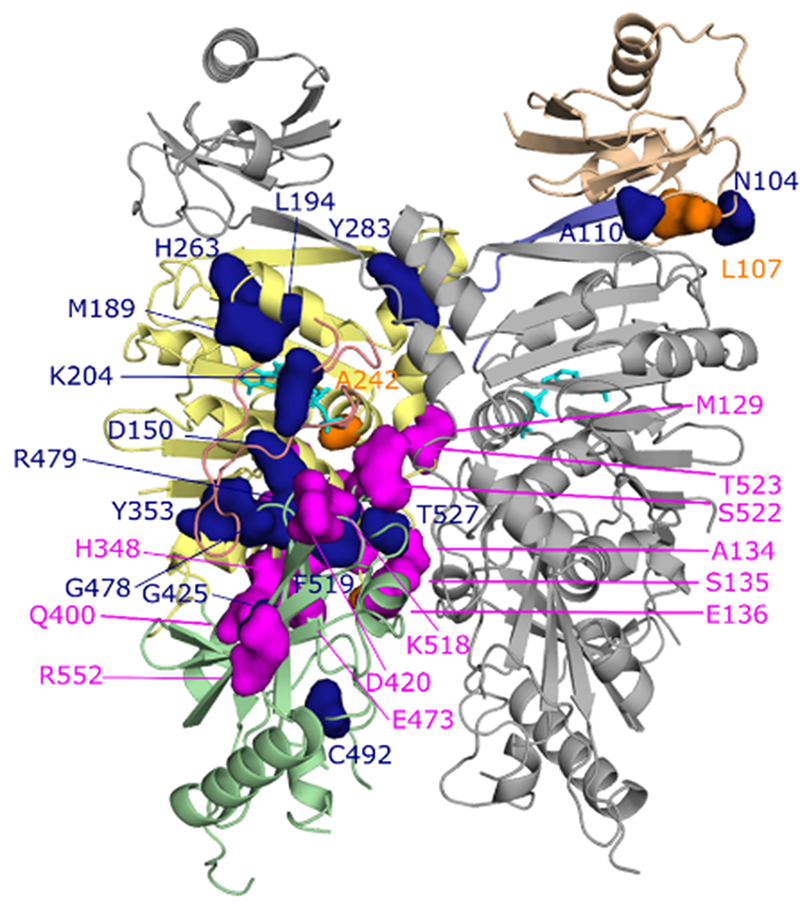
Crystal structure of the GHKL-ATPase domain of human SMCHD1 bound to ATP (cyan) as a functional dimer (one monomer is in all gray). Dimerization is stabilized by a domain-swap of the N-terminal ubiquitin-like (tan) domain with respect to the ATPase (yellow) and transducer (green) domains. This structure reveals the locations of congenital arhinia (magenta) or Facioscapulohumeral Muscular Dystrophy type 2 (FSHD2, blue) disease-associated mutations. Orange residues represent locations of mutations associated with both diseases.
References
- Pedersen, L. C., Inoue, K., Kim, S., Perera, L., and Shaw, N. D. (2019) A ubiquitin-like domain is required for stabilizing the N-terminal ATPase module of human SMCHD1. Commun Biol 2, 255


