Defining Gene Regulatory Mechanisms

Paul A. Wade, Ph.D.
Research Summary
Paul A. Wade, Ph.D., is Chief of the Epigenetics and RNA Biology Laboratory and head of the Eukaryotic Transcriptional Regulation Group. The long-term goals of the laboratory are to understand how eukaryotic cells regulate genes within the context of chromatin (popularly termed epigenetics), how such events relate to biological properties of cells, and how environmental cues impact regulation. The laboratory has a special interest in cancer; studies within the group focus on chromatin-dependent regulation of transcriptional networks that influence risk of cancer development or dictate disease-relevant properties of cancer cells.
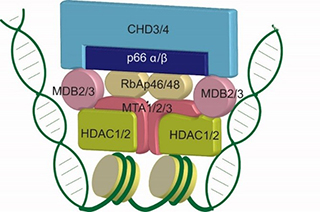
A major interest of the laboratory is the nuclear enzyme Mi-2/NuRD complex. This complex is composed of 6 core subunits each of which is encoded by 2 or more gene paralogs. Combinatorial assembly of these paralogs results in the formation of multiple complexes with both common and unique functions. Mi-2/NuRD complex carries two distinct enzymatic activities (ATP-dependent chromatin remodeling and lysine deacetylation) in a machine that also possesses multiple domains that read DNA and histone modification status. The laboratory is pursuing biochemical reconstitution of the complex from recombinant subunits, determination of complex localization using genomics (Shimbo et al., PLoS Genetics, 2013), and molecular and genetic analyses of Mi-2/NuRD influence on local chromatin properties and gene transcription. Interestingly, Mi-2/NuRD complex subunits have recently been described as targets of mutation in human diseases including cancer and developmental diseases (Snijders Blok et al., Nature Communications, 2018). The laboratory is actively pursuing biochemical and molecular genetic systems to model the impact of human disease-causing mutations in NuRD subunits on chromatin remodeling and gene regulation.
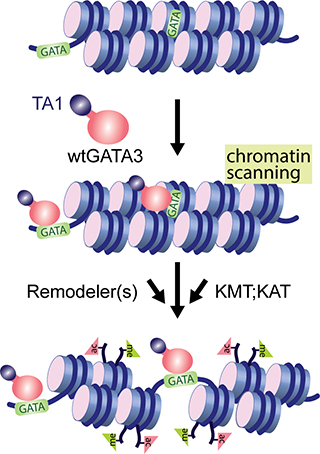
Packaging of eukaryotic DNA into chromatin forms a physical barrier to proteins, such as transcription factors, that need access to DNA to read sequence. Current models suggest that a subset of transcription factors can penetrate the chromatin barrier to productively interact with their cognate recognition sequences, ultimately resulting in formation of a cis-regulatory module (also known as an enhancer). The mechanistic details permitting some transcription factors to ‘pioneer’ local alterations in chromatin properties are poorly understood but central to development, to disease, and to cellular responses to the environment. Our laboratory is investigating ‘pioneer’ activity of a model transcription factor, GATA3 (Takaku et al., Genome Biol, 2016), which is required for mammary gland development and is integral to the transcriptional program found in the most abundant molecular subtype of human breast cancer. Interestingly, recent genomic analyses of breast tumors indicate that GATA3 is a frequent target of cancer-specific mutation. Ongoing studies in the laboratory seek to understand the fundamental properties of GATA3-dependent gene regulation and how these are altered by mutation in cancer (Takaku et al., Nature Communications, 2018).
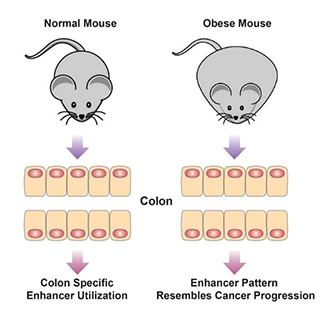
One mechanism by which environmental cues influence biological outcomes is alteration of the epigenome. Diet constitutes an important means by which animals interact with their environment, with powerful downstream impacts on health. The laboratory has developed an ongoing interest in diet and obesity and how these environmental inputs influence health. We employ an animal model of diet-induced obesity that permits direct assessment of how diet and obesity impact chromatin features in colon and liver (Li et al., Cell Metabolism, 2014; Li et al., Cell Reports, 2018; Qin et al., Genome Biol, 2018). These studies have indicated that an obesogenic diet high in fat and low in fermentable sources of fiber results in alterations in the epigenome and transcriptional network in both the colon and liver. Systems level analysis of these outcomes predicts that the altered transcriptional program and signaling network in obese mice increases risk of development of colorectal cancer.
Major areas of research:
- Chromatin remodeling, epigenetics and transcriptional networks
- Pioneer transcription factors
- Diet, obesity, metabolism and microbiome and their relationship to disease risk
Current projects:
- Structure/function analysis of Mi-2/NuRD complex
- Characterization of the influence of local chromatin features and chromatin remodeling/modification on pioneer transcription factors
- Obesity and metabolism and their impact on chromatin features, gene regulation and disease risk
Wade received his Ph.D. degree in Molecular Biology from Indiana University in 1994 and trained as a postdoctoral fellow in the laboratory of Alan Wolffe at NICHD. Following 4 years as an Assistant Professor in the Department of Pathology and Laboratory Medicine, Emory University School of Medicine, he came to NIEHS in 2004. He served as Director of the Epigenomics Core Facility from its inception in 2009 to 2016 and also served as Acting Deputy Scientific Director from 2015 to 2017.


