Features
From Cells to Solutions: Emerging Tools for Studying Health and Disease
The NIEHS Superfund Research Program (SRP) hosted a Risk e-Learning webinar series focused on how grant recipients are using innovative, human-relevant technologies to better characterize the biological effects of chemicals. New Approach Methodologies (NAMS), including advanced cell-based assays, organoids, and computational modeling approaches, are expanding the toolbox researchers use to answer previously difficult or unanswerable questions. Together with representatives from across NIH and EPA, SRP-funded researchers discussed how these emerging methodologies are being applied to uncover mechanistic insights, improve predictive accuracy for human health outcomes, and refine risk assessment frameworks.
Multi-Cellular Systems, Modeling, and Simulations to Advance Environmental Health Research
Advances in cell-based systems, modeling, and simulations are improving researchers’ understanding of complex environmental health topics, including how chemicals interact inside the body, their toxicity, and how they may cause disease.

Researchers from the Michigan State University (MSU) SRP Center developed a “plug and play” multicellular in-vitro system that can mimic interactions between different cells during human development. The model incorporates liver and thyroid cells to help researchers investigate how different organ systems interact with one another when exposed to contaminants, better representing what happens in the body.
Led by Brian Johnson, Ph.D., the team is using this model to map out how different chemicals like PFAS and dioxins affect cellular pathways that can lead to thyroid disruption. The researchers then developed a unique computational approach to predict how an organism might respond to a hazardous exposure based on data gleaned from their model. According to the research team, combining this multicellular system and computational approaches may help predict compounds that contribute to thyroid disruption and serve as a blueprint for future NAMs.
Scientists at the University of North Carolina at Chapel Hill (UNC) SRP Center are using NAMs to investigate how different environmental exposures may affect different organ systems. One project, led by Rebecca Fry, Ph.D., director of the UNC SRP Center, used placental cells from patients to examine how exposure to chemicals like PFAS affect fetal development and placental function. The scientists found that PFAS accumulated in placental cells and may disrupt placental migration, a critical process during fetal development. In a similar study, UNC researchers exposed placental cells to arsenic and cadmium to uncover how these exposures affect genes in ways that may harm placental function.

Researchers at the University of Arizona SRP Center are using NAMs to understand how realistic environmental conditions affect the bioaccessibility of arsenic in particulate matter from mining waste. Bioaccessibility refers to the fraction of a contaminant that has the potential to be absorbed into the bloodstream — a key determinant of potential toxicity. Specifically, they are looking at how weathering changes the geologic makeup and chemical properties of arsenic bound to particulate matter, and how those changes influence bioaccessibility.
Led by Jon Chorover, Ph.D., the team created a simulated gastrointestinal tract in the lab to model digestion of this particulate matter under different conditions. The scientists found that the bioavailability of arsenic from particulate matter varies widely and depends on characteristics like the type of arsenic and the co-occurrence of other contaminants.
3D Models and Technologies to Illuminate Biological Effects of Contaminants
Advancements in 3D models and other technologies are enabling scientists to better understand how contaminants contribute to human diseases, including harming DNA, altering placental function, and contributing to asthma.
Led by Susan Tilton, Ph.D., researchers from the Oregon State University (OSU) SRP Center are using newly-developed 3D lung cell models to answer questions about the respiratory toxicity of polycyclic aromatic hydrocarbons (PAHs) — a family of chemicals that can contaminate air, water, and soil. Airway epithelial cells are the lung’s first line of defense against PAHs, and these cells communicate with macrophage immune cells to trigger inflammation after exposure. By combining both cell types at a unique air-liquid interface, the new cell model better mimics conditions within the lungs, making it easier for researchers to understand and predict how pollutants like PAHs affect respiratory health. The 3D model can also be modified to mimic pre-existing human respiratory disease to better clarify how having asthma, for example, may increase susceptibility to PAH exposure.
OSU SRP scientists also collaborated with the Pacific Northwest National Laboratory to develop a physiologically based pharmacokinetic model — a common toxicology approach used to understand how a substance moves and changes in the body. According to the researchers, combining these two approaches can help scientists track how PAHs are metabolized, measure DNA damage, and identify harmful health effects based on dose.

The organ-on-a-chip (OOC) models developed at TAMU capture the complexity of different cells involved in the fetal membrane and placenta, respectively, indicated by different colors here. (Image courtesy of the TAMU SRP Center)
Researchers at the Texas A&M University (TAMU) SRP Center are investigating how PFAS exposures may contribute to preterm birth using organ-on-a-chip technologies. Led by Arum Han, Ph.D., the scientists developed an organ-on-a-chip that mimics the unique membranes and placental cells at the interface between a mother and a developing fetus. This approach allows them to monitor the transport, cellular response, and inflammatory effects of PFAS compounds, and revealed that fetal membrane cells are more reactive and susceptible to inflammatory responses than placental cells. The chips also help researchers identify the underlying mechanisms by which PFAS can harm human health.
To enable more comprehensive testing for PFAS and complex chemical mixtures, the scientists are exploring ways to increase the automation and throughput of the chip studies. According to the team, the organ-on-a-chip design shows promising potential for prioritizing chemicals for further study and informing health-protective decisions for sensitive populations, such as pregnant women and unborn children.
Innovative Methods for Understanding Chemical Toxicity
To provide faster, more accurate data on chemical toxicity, scientists are moving beyond traditional lab tests. These approaches use human-relevant models — including three-dimensional (3D) organoids and sophisticated computer modeling — to understand the precise dose at which chemicals trigger biological harm and the specific mechanisms behind it.
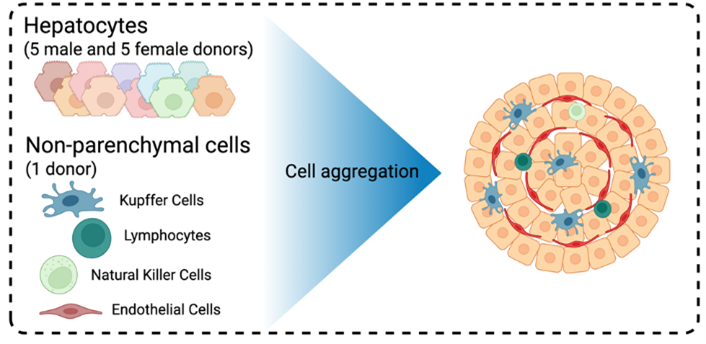
Researchers at the University of Southern California (USC) SRP Center are using 3D human liver spheroids to characterize how PFAS mixtures affect liver metabolism and disease. By creating their spheroids from the liver cells of multiple human donors, the researchers can capture a wide range of human genetic diversity, providing a more realistic picture of how different people might respond to chemical exposures. The 3D spheroid model mimics the architecture and cell-to-cell interactions of the human liver, overcoming limitations of traditional 2D cultures.
Led by Vaia Lida Chatzi, Ph.D., and Ana Maretti Garcia, Ph.D., the USC team is using a combination of transcriptomics, epigenomics, proteomics, and other methods to identify key changes following PFAS exposure, such as increased lipid accumulation and the upregulation of cancer-related pathways. By integrating data from these spheroids with human exposure data, they identified biomarkers and mechanistic pathways that link PFAS exposure to metabolic dysfunction and illuminated molecular mechanisms of PFAS-induced liver injury.
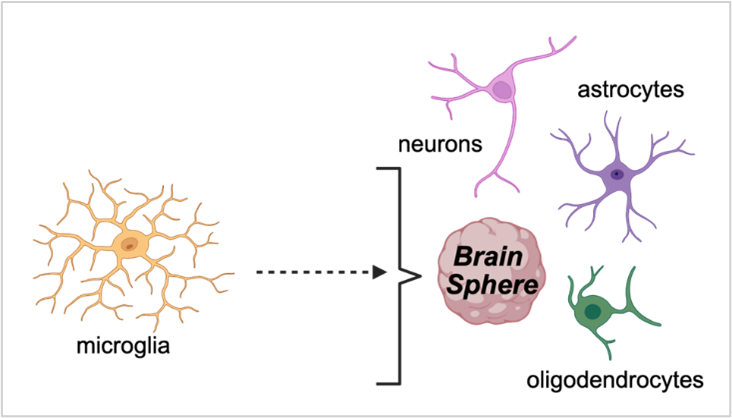
Scientists at the Duke University SRP Center are using 3D brain spheres, derived from human-induced pluripotent stem cells (iPSC), to study the effects of chemical co-exposures on developmental neurotoxicity. Their model overcomes the challenges of traditional lab models to study neurotoxicity because the spheres contain a variety of interacting brain cells — including neurons, astrocytes, and microglia — and exhibit spontaneous electrical activity, which allows for functional neurotoxicity assessments. This system can capture cell-specific effects, such as selective neuron loss. The model detects dose-dependent molecular and functional changes even before obvious signs of cell damage are visible, revealing biologically relevant safety thresholds for developmental neurotoxicity.
The team, including Duke researcher Susan Murphy, Ph.D., and trainee Guru Ulaganathan, is using temporal modeling to assess how early versus late exposures affect brain development. They are also incorporating nervous system immune cells to study immune-mediated neurotoxicity.
The TAMU SRP Center is developing a suite of NAMs to rapidly characterize hazards and risks from exposure to chemicals and mixtures in the aftermath of environmental disasters. To determine the potential health risks posed by complex environmental mixtures, the center uses in vitro hazard identification. For example, following Hurricane Harvey, researchers collected sediment samples from flooded areas and tested extracts of these real-world mixtures directly on human iPSC-derived cell models. This approach allows scientists to identify areas of greatest concern based on the mixture’s total biological activity to prioritize cleanup efforts.
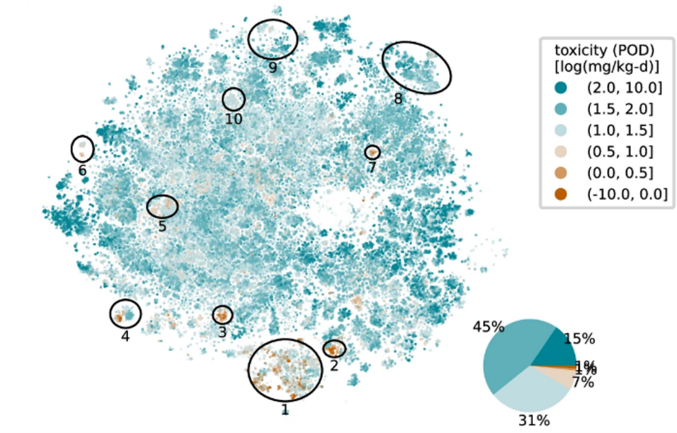
To address the vast number of chemicals that lack any safety data, the TAMU team, led by Weihsueh Chiu, Ph.D., created computer-based dose-response tools. One method uses a massive database of existing animal studies to generate reliable "surrogate" safety levels for over 10,000 chemicals. For chemicals with no animal data at all, they developed a two-stage machine learning model that predicts a chemical's point of departure (POD) — the lowest dose that triggers a biological response — based on its structure alone. This computational approach expands researchers’ ability to evaluate chemicals for potential health risks.
Other Relevant SRP-Funded Research
In addition to the speakers featured in the webinar series, SRP grant recipients across the country continue to develop and implement research tools to complement more traditional research approaches, including specialized cell models, high throughput approaches, and machine learning and in silico methods. For example:
- Naomi Halas, Ph.D., and colleagues at the Baylor College of Medicine SRP Center are using machine learning techniques to better detect and identify specific PAHs in human placenta tissue. This approach will improve exposure characterization and public health surveillance.
- Researchers at the Duke SRP Center, led by Nishad Jayasundara, Ph.D., developed a statistical model to estimate the neurobehavioral effects of metal and metal mixtures in drinking water. Their approach offers a unique method to analyze complex mixtures data and identify “bad actors” driving adverse effects.
- April Z. Gu, Ph.D., and collaborators at the Northeastern University SRP Center developed a new computational approach that uses machine learning and advanced algorithms to link biological changes from high throughput cell studies with health outcomes observed in animal studies to classify and predict chemicals that cause harm. According to the team, their approach can help researchers reduce the time, cost, and complexity of screening chemicals for further study.
- Researchers at the North Carolina State University SRP Center developed a new approach that combines advanced imaging with machine learning to rapidly screen for changes to immune cells. Their approach can be adapted for other cell types in a variety of organisms.
- At the University of Kentucky SRP Center, researchers are using machine learning techniques to interpret and predict how compounds are processed, or metabolized, in the body. This information is important for protecting human health because sometimes metabolites of chemicals are more toxic than the original compound.
to Top



