About EagleView
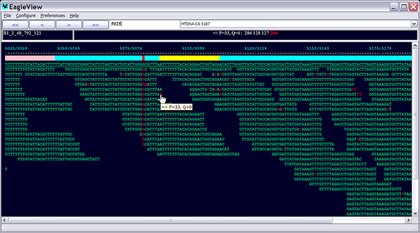
EagleView is an information-rich viewer for next-generation genome assembles with data integration capability. EagleView can display a dozen different types of information including base qualities, machine specific trace signals, and genome feature annotations. It provides an easy way for inspecting visually the quality of a genome assembly and validating polymorphism candidate sites (e.g., SNPs) reported by polymorphism discovery tools. It can also facilitate data interpretation and hypothesis generation. EagleView is a multi-platform application developed with C++ and is available for all three major platforms: Windows, Linux, and Mac OS.
Key Features
- fast and memory efficient
- genome annotation integration
- pin-point view of machine-specific signals
- pin-point view of base quality
- pin-point view of read id and strand
- distinct mark for discrepancy sites
- navigation by read id and contig id
- data utility tools included
- navigation by genomic features or user defined locations
- zooming and printing capability
- navigation by both unpadded and padded positions
- customizable font and color for viewer
Availability
EagleView is freely available to the public, and the latest version (v2.2) can be downloaded at the following links.
Linux OS
- EagleView for 64-bit Debian-based Linux systems (e.g. Ubuntu) - EagleView_v2.2_Linux64.tar.gz (6MB)
- EagleView for 32-bit Debian-based Linux systems (e.g. Debian) - EagleView_v2.2_Linux32.tar.gz (6MB)
- EagleView for 64-bit Redhat-based Linux systems (e.g. CentOS) - EagleView_v2.2_CentOS5_x64.tar.gz (6MB)
MS Windows
- EagleView for 64-bit MS Windows - EagleView_v2.2_Win64.exe (2MB)
- EagleView for 32-bit MS Windows - EagleView_v2.2_Win32.exe (1MB)
Mac OS X
- EagleView for MacOS X (with intel CPU) - EagleView_v2.1_MacOSX.dmg (3MB)
EagleView should be cited as
Weichun Huang and Gabor T Marth, EagleView: a genome assembly viewer for next-generation sequencing technologies, Genome Res.,June 11, 2008, 10.1101/gr.076067.108
Installation instruction
MS Windows Distributions
- Package: A single binary installer for either 32 or 64-bit system (XP or Vista).
- Installation: Double click EagleView installer and simply follow its instructions.
- Launch: EagleView can launch from either Windows program menu or its shortcut at your Desktop
Linux Distributions
- Package: EagleView is distributed as a tar.gz package for both 32-bit and 64-bit Linux systems.
- Installation: Simply extract EagleView tar.gz by the following command: tar xvfz EagleView-*.tar.gz
- Launch: Enter your extraction directory, and issue the following command at your Linux terminal window: ./bin/EagleView &
Apple Mac Distribution
- Package: EagleView Mac version is available for Mac OS X 10.4 or later with Intel CPU. The distribution is a disk image containing Mac OS X installation Package
- Installation: You need to have the administrator privileges to install EagleView. Installation is done by double clicking EagleView Installer the disk image
- Launch: Double click EagleView application at /Applications/eagleview. The data tools for EagleView should be used in the terminal window.
Datasets for Examples and Tests
Small example datasets


